- R&D
- Clinical Diagnosis
- News
- Company
About Us
Quick Biology is a CLIA-certified, high-throughput genomic center equipped with all majornext-generation sequencing. QB provides state-of-the-art genomics technologies,comprehensive services, specialized expertise enabling these services in a cost-effectiveand timely manner to serve basic science and translational/clinical research, QB's serviceincluded, but not limited to, RNA sequencing, whole exome sequencing, whole genomesequencingATAC sequencing and single cell RNA sequencing.





Why Quickbiology
Proprietary NGS Technologies
As Celemics' exclusive technologies for designing probes and optimizing assay and chemistry formarket-leading performance
Panel Diversity
Wide range of validated ready-to-use panels with flexiblecustomization option, also applicable tovarious challenging sample type
Client-oriented Customization Service
End-to-end customization and assay development service,including support for regulatory affairs
Satisfied Customers
Perception Enabled
Quest can assist you in solving any upcoming IT challengesallowing you to quickly enter the production phase and
achieve your next IT goal in a short period of time, and it will
continue to do so in the future. Control your enterprise on
multiple operating systems and devices through
comprehensive software and hardware inventory and asset
management functions. Whether it's internal.....

Equiped with the Latest Sequencing Instruments

10x Chromium Controller

Illumina NovaSeg 6000

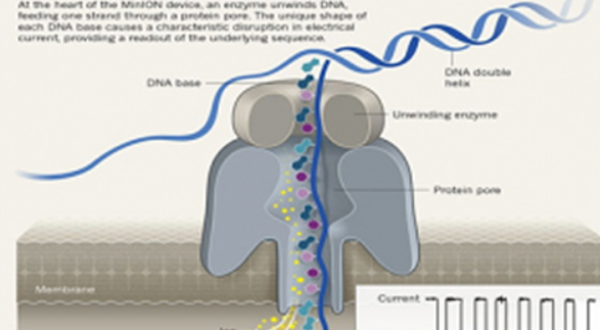
Nanopore

Illumina Novaseg X